MerlinViewer
Visualising Merlin output data
During the development of mapping software it is important to valid the program using simple pedigree structures which can be analysed using other software packages such as Merlin. However, when analysing high density SNP chip genotype data with Merlin, the output consists of a very large results table which due to its size may be difficult to process by eye. To overcome this problem MerlinViewer was developed to visiualise the data as a scatter graph of the LOD, HLOD or Alpha score against chromosomal position.
The MerlinViewer program first reads the appropriate map file and then the Merlin output text file. After pressing the 'View' button the data is read and displayed as a scatter graph with the x-axis representing the SNPs position and the y-axis representing the SNPs linkage score.
The data underlying a region of interest can be exported either to a web page or to a tab-delimited text file. To select such a region, place the mouse cursor at the start of the region and while holding down the left mouse button, drag the cursor to the end of the region. The currently selected region will be delimited by two black vertical lines (Figure 1). To save the data, press and enter the name of the output file and filename extension.
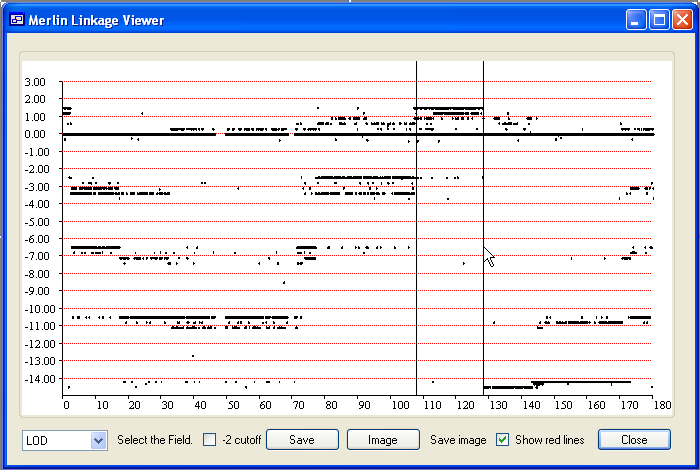
Figure 1: Chromosome 5 data for a family affected by a dominant APC mutation
Each *.map and Merlin data file must contain the
data for a single chromosome and be the standard format as exported by the
program. MerlinViewer and a set of map and results files can be downloaded
here
Note: this program requires the .NET framework version 2.0 to be installed
(available through Windows Update).
- Merlin details
- Merlin was developed by G.R. Abecasis at the Center for Statistical Genetics, University of Michigan and has its own web page here
- Merlin citation
- Abecasis GR, Cherny SS, Cookson WO and Cardon LR (2002).
Merlin-rapid analysis of dense genetic maps using sparse gene flow trees.
Nature Genetics 30:97-101.